Professor
Tier 1 Canada Research Chair in plant immunity and Sclerotinia biology
Academic History
- BSc. Genetics and Genetic Engineering, Fudan University (1989)
- Ph.D. Plant Pathology, Oklahoma State Univ., (1995)
- Postdoctoral Fellow, Duke Univ., (1996-1999)
- Scientist, Maxygen. Inc. (1999-2001)
Contact Information
- xinli@msl.ubc.ca
- 604-822-3155
- Office Room 317/Lab Room 377 MSL
- Lab Phone: 604-822-3205
Research Interests
The long-term goal of our research program is to understand the molecular mechanisms of plant innate immunity. We study plant defense against pathogen infection in the context of gene regulation, protein-protein interaction and signal transduction using the model organism Arabidopsis thaliana. Our program aims to discover new regulatory components of plant disease resistance and to elucidate the biochemical functions of a number of regulators we have already identified. Understanding the innate ability of plants to defend themselves against pathogen infection promises to revolutionize disease control practices in our fields using environmentally friendly strategies.
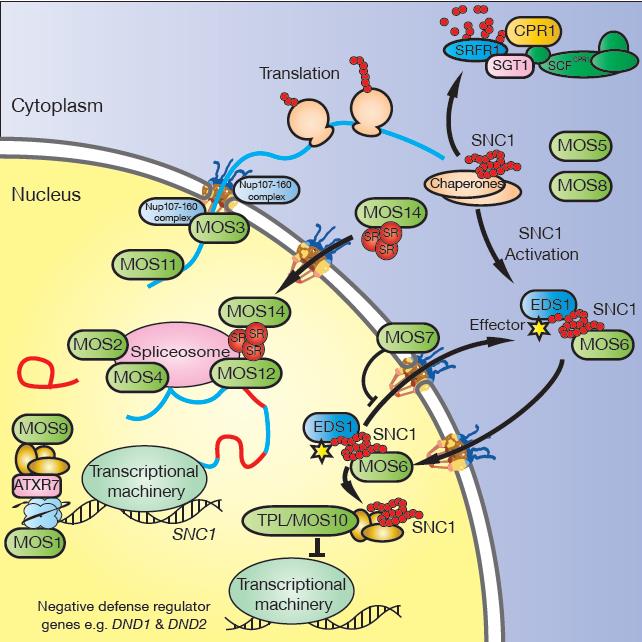
A model depicting the involvement of the MOS proteins in R protein-mediated defense signaling pathways in Arabidopsis, using SNC1 as an example of the journey of TIR-NB-LRR proteins. 1. At the chromosomal level, MOS1, ATXR7, and MOS9 up-regulate the transcription of SNC1 through chromatin remodeling. 2. MOS2, MOS4, and MOS12 are required for proper splicing of the transcripts of SNC1. 3. The Nup107-160 complex and MOS11 have key roles in the export of total mRNA (including mature mRNA of SNC1), which is required for effective defense. 4. MOS5 is an E1 ubiquitin-activating enzyme, an essential component of the ubiquitination cascade, required for the regulation of defense signaling components. As an example, the SCFCPR1 E3 ubiquitin ligase complex targets SNC1 for degradation, which prevents autoimmunity caused by overaccumulation of R proteins. MOS8 positively regulates plant defense, possibly through prenylation, which affects the targeting of defense regulators. 5. MOS6 and MOS7 are involved in the nucleocytoplasmic shuttling of defense signaling molecules such as SNC1, EDS1, and NPR1. As with RPS4, EDS1 is probably required for the nuclear localization and activation of SNC1 upon recognition of its corresponding effector (Bhattacharjee et al. 2011; Heidrich et al. 2011). MOS14 is required for the nuclear import of splicing factors, which may affect defense regulator RNA processing. 6. MOS10 activates the SNC1-mediated defense through transcriptional repression of negative regulators of defense such as DND1 and DND2.
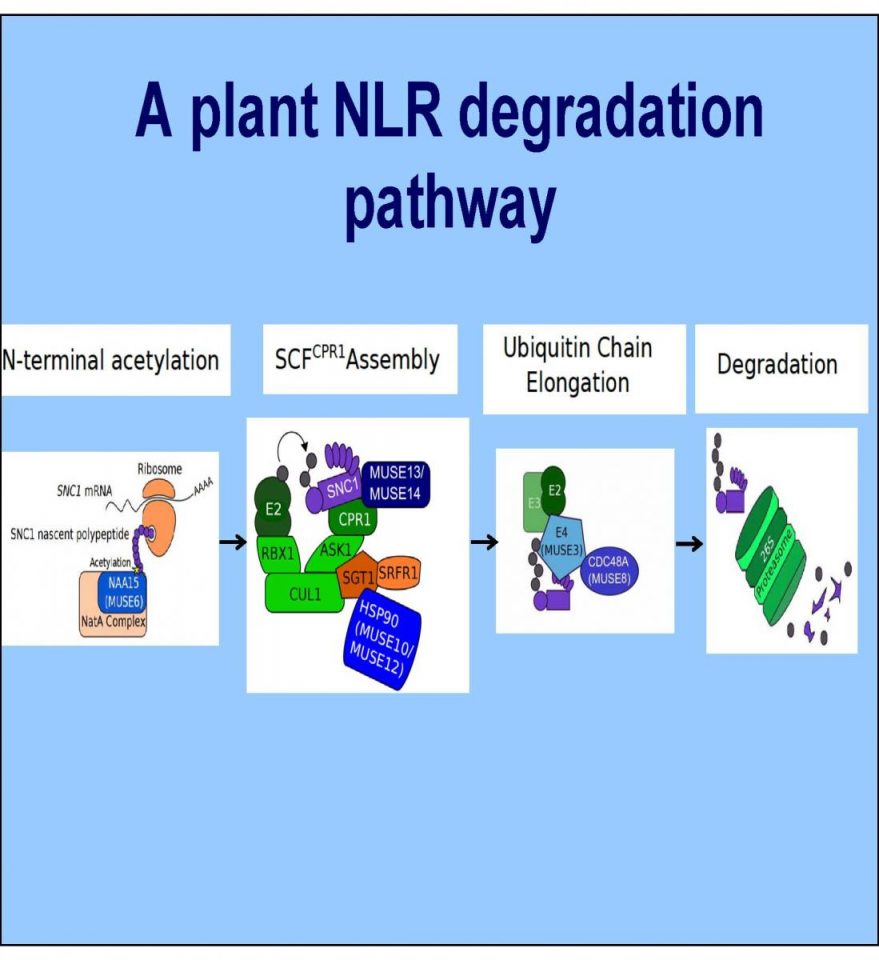
The Li lab also investigates the biology of fungal pathogens including Sclerotinia sclerotiorum and Botrytis cinerea. These pathogens cause severe damages in agriculture worldwide, and understand their basic developmental and virulence processes will be essential for designing sustainable methods for crop protection.

Models comparing the status of chromosomes in the nuclei of S. sclerotiorum ascospores (A) and B. cinerea conidia (B). (A) In the classic model, the two nuclei in each S. sclerotiorum ascospore contain identical complement of 1N chromosomes (1N=16). In our new model, the 16 chromosomes are segregated into two nuclei in an unfixed manner. (B) In the classic model, each nucleus in the B. cinerea conidium contains a full complement of 1N chromosomes (1N=18). In our new model, the 18 chromosomes are sorted into different nuclei inside each conidium in an unfixed manner.

Team Members
If you are interested in pursuing a PhD degree in plant immunity and/or Sclerotinia biology, please contact Dr. Xin Li (xinli@msl.ubc.ca).
Postdoctoral Fellow:
Dr. Weijie Huang (2023-)
Dr. Ning Cui (2023-)
Ph.D. Students:
Xueru Liu (2020- )
Tina Tan (2021-)
Edan Jackson (2023-)
Zhengxi Gong (2024-)
Ruonan He (2024-)
Ziyang Xiao (2025-)
MSc Students:
Thilini Weerasinghe (2023-)
Lois Chen (2024-)
Josh Li (2023-)
Lab Alumni:
Dr. Marcel Wiermer, Ph.D (2005)
Dr. Sandra Goritschnig, Ph.D. (2006)
Dr. Hugo Germain, Ph.D. Plant Science (2007)
Dr. Kristoffer Palma, Ph.D. Genetics (2007)
Dr. Jacqueline Monaghan, Ph.D (2010)
Patrick Gannon, MSc, Botany (2011)
Dr. Tabea Weihmann, Ph.D (2011)
Virginia Woloshen, MSc, Botany (2012)
Dr. Yu Ti Cheng, Ph.D (2013)
Dr. Yan Huang, Ph.D (2013)
Chipan Zhu, MSc, Botany (2014)
Dr. Fang Xu, Ph.D (2014)
Dr. Kaeli Johnson, Ph.D (2016)
Dr. Oliver Xiaoou Dong, Ph.D (2016)
Dr. Shuai Huang, Ph.D (2016)
Dr. Meixuezi Tong, Ph.D (2016)
Dr. Charles Copeland, PhD (2018)
Dr. Jianhua Huang, PhD (2019)
Dr. Zhongshou Wu (2015- 2021)
Dr. Wanwan Liang (2015-2020)
Dr. Lei Tian (2019-2023)
Dr. Paul Kapos (2014-2020)
Dr. Kevin Ao (2017-2023)
Rowan van Wersch (2016-2024)
Dr. Yan Xu (2017-2023)
Dr. Karen Thulasi Devendrakumar (2017- 2024)
Dr. Tongjun Sun (2018- 2021)
Nanbing Zhang (2018- 2020)
Solveig van Wersch (2018- 2020)
Selected Publications
Zhang, Y., Goritschnig, S., Dong, X., and Li, X. 2003. A gain-of-function mutation in a plant disease resistance gene leads to constitutive activation of downstream signal transduction pathways in the snc1 mutant. Plant Cell 15 (11): 2636-46.
Zhang Y. and Li X. 2005. A putative Nucleoporin 96, is required for both basal defense and constitutive resistance responses mediated by snc1. Plant Cell 17: 1306-16.
Palma K, Zhang Y, and Li X. 2005. An Importin alpha homolog, MOS6, plays an important role in plant innate immunity. Current Biology 15(12):1129-35.
Zhang Y.,Cheng, Y.T., Bi, D., Palma K., and Li, X. 2005. MOS2, a protein containing G-patch and KOW motifs, is essential for innate immunity in Arabidopsis thaliana. Current Biology 15(21):1936-42.
Goritschnig S, Zhang Y, and Li, X. 2007. The ubiquitin pathway is required for innate immunity in Arabidopsis. Plant Journal 49(3):540-51.
Palma K., Zhao Q., Cheng Y.T., Bi D., Monaghan J., Cheng W., Zhang Y., and Li X. 2007. Regulation of plant innate immunity by three proteins in a complex conserved across the plant and animal Kingdoms. Genes & Development 15; 21(12):1484-93.
Monaghan J., Xu F., Zhao Q., Palma K., Chen S., Zhang Y. and Li X. 2009. Two Prp19-like U-box proteins in the MOS4-Associated Complex play redundant roles in plant innate immunity. PLoS Pathogens, Jul;5(7):e1000526.
Cheng Y.T., Germain H., Wiermer M., Bi D., Garcia A.V., Wirthmueller L., Despres C., Parker JE, Zhang Y., and Li X. 2009. MOS7 is required for plant innate immunity and nuclear accumulation of defense regulators. Plant Cell 21(8): 2503-16.
Germain H., Qu N., Cheng Y.T., Lee E.K., Huang H., Dong O.X., Gannon P., Huang S., Ding P., Li Y., Sack F., Zhang Y., and Li X. 2010. MOS11: a new component in the mRNA export pathway. PLoS Genetics 6 (12) e1001250.
Bi D, Johnson KC, Zhu Z, Huang Y, Chen F, Zhang Y, Li X. 2011. Mutations in an Atypical TIR-NB-LRR-LIM Resistance Protein Confer Autoimmunity. Front Plant Sci. 2:71. doi: 10.3389/fpls.2011.00071.
Cheng Y.T., Li Y., Huang S., Huang Y., Dong X., Zhang Y. and Li X. 2011. Stability of plant immune-receptor resistance proteins is controlled by SKP1-CULLIN1-F-box (SCF)-mediated protein degradation. PNAS 108(35):14694-9.
Xu F., Xu S., Wiermer M., Zhang Y. and Li X. 2012. The cyclin L homolog MOS12 and the MOS4-associated complex are required for proper splicing of plant resistance genes. Plant Journal, 70(6):916-28.
Cheng Y.T. and Li X. 2012. Ubiquitination in NB-LRR-mediated immunity. Current Opinion in Plant Biology 15(4):392-99.
Johnson K.C., Dong O.X., Huang Y., Li X. 2013. A Rolling Stone Gathers No Moss, but Resistant Plants Must Gather Their MOSes. Cold Spring Harb Symp Quant Biol. Feb 21.10.1101/sqb.2013.77.014738
Huang Y., Chen X., Liu Y., Rothe C., Copeland C., McFarlane H.E., Huang S., Lipka V., Wiermer M. and Li X. 2013. Mitochondrial AtPAM16 is required for plant survival and negative regulation of plant immunity. Nature Communications, 4:2558. doi: 10.1038/ncomms3558.
Huang Y., Minaker S., Roth C., Hieter P., Lipka V., Wiermer M. and Li X. 2014. An E4 ligase facilitates polyubiquitination of plant immune receptor resistance proteins. Plant Cell 26(1):485-96.
Huang S., Monaghan J., Zhong X., Lin L., Sun T., Dong O.X. and Li X. 2014. HSP90s are required for NLR immune receptor accumulation in Arabidopsis. Plant Journal 79(3):427-39.
Xu F., Kapos P., Cheng Y.T., Li M., Zhang Y. and Li X. 2014. NLR-Associating Transcription Factor bHLH84 and Its Paralogs Function Redundantly in Plant Immunity. PLoS Pathog. 10(8):e1004312. doi: 10.1371/journal.ppat.1004312.
Xu F., Cheng Y.T., Kapos P., Huang Y. and Li X. 2014. P-Loop-Dependent NLR SNC1 Can Oligomerize and Activate Immunity in the Nucleus. Mol Plant 7(12) 1801-1804.
Xu F., Zhu C., Cevik V., Johnson K., Liu Y., Sohn K., Jones J.D., Holub E.B. and Li X. 2015. Autoimmunity conferred by chs3-2D relies on CSA1, its adjacent TNL-encoding neighbour. Sci Rep. 5:8792. doi: 10.1038/srep08792.
Li X., Kapos P. and Zhang Y.. NLRs in plants. 2015. Curr Opin Immunol. 32C:114-121. doi: 10.1016/j.coi.2015.01.014.
Xu F., Huang Y., Li L., Gannon P., Linster E., Huber M., Kapos P., Bienvenut W., Polevoda B., Meinnel T., Hell R., Giglione C., Zhang Y., Wirtz M., Chen S. and Li X. 2015. Two N-Terminal Acetyltransferases Antagonistically Regulate the Stability of a Nod-Like Receptor in Arabidopsis. Plant Cell 27(5):1547-62.
Huang S., Chen X., Zhong X., Li M., Ao K., Huang J. and Li X. 2016. Two redundant plant TRAF proteins participate in NLR immune receptor turnover. Cell Host & Microbe 19(2):204-15.
Li X. and Zhang Y. Suppressor Screens in Arabidopsis. Methods Mol Biol. 2016;1363:1-8. doi: 10.1007/978-1-4939-3115-6_1.
Dong O.X., Tong M., Bonardi V., Kasmi F.E., Woloshen V., Wünsch L.K., Dangl J.L. and Li X. TNL-mediated immunity in Arabidopsis requires complex transcriptional regulation of the redundant ADR1 gene family. New Phytologist 210(3):960-973.
Johnson K.C.M., Yu Y., Gao L., Eng R.C., Wasteneys G.O., Chen X. and Li X. 2016. A partial loss-of-function mutation in an Arabidopsis RNA polymerase III subunit leads to pleiotropic defects. Journal of Experimental Botany 67(8):2219-30.
Copeland C., Woloshen V., Huang Y. and Li X. 2016. AtCDC48A is involved in the turnover of an NLR immune receptor. Plant J. 2016 Jun 24. doi: 10.1111/tpj.13251. [Epub ahead of print]
Huang S., Balgi A., Pan Y., Li M., Zhang X., Du L., Zhou M., Roberge M. and Li X. 2016. Identification of methylosome components as negative regulators of plant immunity using chemical genetics. Mol Plant. 9(12):1620-1633.
Johnson KC, Zhao J, Wu Z, Roth C, Lipka V, Wiermer M, and Li X. 2017. The putative kinase substrate MUSE7 negatively impacts the accumulation of NLR proteins. Plant J. 89(6):1174-1183.
Wu Z., Huang S., Zhang X., Wu D., Xia S. and Li X. 2017. Regulation of plant immune receptor accumulation through translational repression by a glycine-tyrosine-phenylalanine (GYF) domain protein. eLife2017 Mar 31;6. pii: e23684
Tong M, Kotur T, Liang W, Vogelmann K, Kleine T, Leister D, Brieske C, Yang S, Lüdke D, Wiermer M, Zhang Y, Li X, Hoth S. 2017. E3 ligase SAUL1 serves as a positive regulator of PAMP-triggered immunity and its homeostasis is monitored by immune receptor SOC3. New Phytol. 215(4):1516-1532.
Ding Y, Sun T, Ao K, Peng Y, Zhang Y, Li X, Zhang Y. 2018. Opposite Roles of Salicylic Acid Receptors NPR1 and NPR3/NPR4 in Transcriptional Regulation of Plant Immunity. Cell. 2018 Apr 11. pii: S0092-8674(18)30376-3. doi: 10.1016/j.Cell.2018.
Dong OX, Ao K, Xu F, Johnson KCM, Wu Y, Li L, Xia S, Liu Y, Huang Y, Rodriguez E, Chen X, Chen S, Zhang Y, Petersen M, and Li X. 2018. Individual components of paired typical NLR immune receptors are regulated by distinct E3 ligases. Nature Plants, doi: 10.1038/s41477-018-0216-8.
Liang W, van Wersch S, Tong M, and Li X. 2019. TIR-NB-LRR immune receptor SOC3 pairs with truncated TIR-NB protein CHS1 or TN2 to monitor the homeostasis of E3 ligase SAUL1. New Phytol. 221(4):2054-2066. doi: 10.1111/nph.15534.
Huang J, Sun Y, Orduna AR, Jetter R, and Li X. 2019. The Mediator kinase module serves as a positive regulator of salicylic acid accumulation and systemic acquired resistance. Plant J. 98(5):842-852.
Wu Z, Li M, Dong OX, Xia S, Liang W, Bao Y, Wasteneys G, and Li X. 2019. Differential regulation of TNL-mediated immune signaling by redundant helper CNLs. New Phytol. 222(2):938-953. doi: 10.1111/nph.15665.
Zhang Y and Li X. 2019. Salicylic acid: biosynthesis, perception, and contributions to plant immunity. Curr Opin Plant Biol. 50:29-36. doi: 10.1016/j.pbi.2019.02.004.
van Wersch S and Li X. Stronger When Together: Clustering of Plant NLR Disease resistance Genes. Trends in Plant Sci. 2019 Aug;24(8):688-699. doi: 10.1016/j.tplants.2019.05.005. Epub 2019 Jun 29.
Sun T, Huang J, Xu Y, Verma V, Jing J, Sun Y, Orduna AR, Tian H, Huang X, Xia S, Schafer L, Jetter R, Zhang Y and Li X. 2019. Redundant CAMTA transcription factors negatively regulate the biosynthesis of salicylic acid and N-hydroxypipecolic acid by modulating the expression of SARD1 and CBP60g. Molecular Plant Nov 13. pii: S1674-2052(19)30364-8. doi: 10.1016/j.molp.2019.10.016. [Epub ahead of print]
Wu Z., Tong M., Tian L., Zhu C., Liu X., Zhang Y. and Li X. 2020. Plant E3 Ligases SNIPER1 and SNIPER2 Broadly Regulate the Homeostasis of Sensor NLR Immune Receptors. EMBO Journal Jun 18;e104915. doi: 10.15252/embj.2020104915. Online ahead of print.
Liang W, Tong M, Li X. SUSA2 is an F-box protein required for autoimmunity mediated by paired NLRs SOC3-CHS1 and SOC3-TN2. Nature Communications 2020 Oct 15;11(1):5190. doi: 10.1038/s41467-020-19033-z.
Li X, Zhang Y.A structural view of salicylic acid perception. Nature Plants. 2020 Oct;6(10):1197-1198. doi: 10.1038/s41477-020-0741-0.
Tian L and Li X. Enzyme formation by immune receptors. Science, 04 Dec 2020: Vol. 370, Issue 6521, pp. 1163-1164. DOI: 10.1126/science.abf2833
Tian, H., Chen, S., Wu, Z., Ao, K., Yaghmaiean, H., Sun, T., Huang, W., Xu, F., Zhang, Y., Wang, S., Li, X.**, and Zhang, Y.** (2021). Activation of TIR signalling boosts pattern-triggered immunity. Nature, https://doi.org/10.1038/s41586-021-03987-1.
Xu Y, Ao K, Tian L, Qiu Y, Huang X, Liu X, Hoy R, Zhang Y, Rashid KY, Xia S, Li X.2022. A Forward Genetic Screen in Sclerotinia sclerotiorum Revealed the Transcriptional Regulation of Its Sclerotial Melanization Pathway. MPMI 35(3):244-256.
Tian L, Li X. 2022. TIR domains as two-tiered enzymes to activate plant immunity. Cell. 185(13):2208-2209. doi: 10.1016/j.cell.2022.05.025.
Xu Y, Tan J, Lu J, Zhang Y, Li X. 2023. RAS signaling genes can be used as host-induced gene silencing targets to control fungal diseases caused by Sclerotinia sclerotiorum and Botrytis cinerea. Plant Biotechnol J. 22(1):262-277. doi: 10.1111/pbi.14184. Epub 2023 Oct 16.
Tian L, Li J, Xu Y, Qiu Y, Zhang Y, Li X. 2023. A MAP kinase cascade broadly regulates the lifestyle of Sclerotinia sclerotiorum and can be targeted by HIGS for disease control. Plant J. doi: 10.1111/tpj.16606.
Xu Y, Tian L, Tan J, Huang W, Li J, O’Neil N, Hirst M, Hieter P, Zhang Y, Li X. 2025. Distribution of haploid chromosomes into separate nuclei in two pathogenic fungi. Science 388(6748):784-788. doi: 10.1126/science.abn7811. Epub 2025 May 15.






